This web page was produced as an assignment for Genetics 677, and undergraduate course at UW-Madison.
Gene Phylogeny
While looking for the DNA sequence for KCNQ1 it was found that the entire sequence is over 400,000 nucleotides. With a sequence that long the use of online programs such as Phylogeny and ClustalW were off limits as their maximum sequence size was around 40,000 nucleotides. That being said phylogenetic trees using Treefam and Phylogeny could not be used to construct phlyogenetics trees based on the DNA sequences between the eight homologous species that were found using Homologene.
Phylogenetic Tree
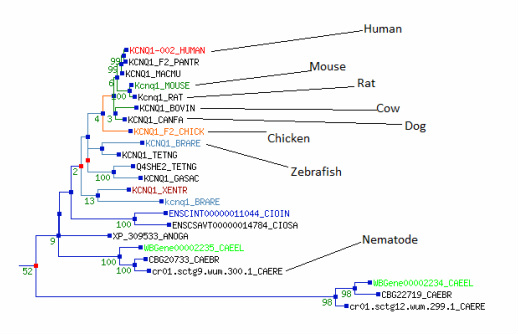
The picture to the left is just part of a phylogenetic tree for the KCNQ gene family. Everything but the part pertaining KCNQ1 has been cropped out. The full tree can be viewed at Treefam. Although there are other species within the tree the eight that I have selected to use thoughout my research have been pointed out. It should be noted that as mentioned before this tree was found within the Treefam database and was not created using the sequences that I found searching through Homologene.
Analysis
Because the DNA sequences were too long Treefam was the only online resource that could be used. This was not because it allowed larger DNA sequences to be inputted but because it allows you to search for phylogenetic information based on a gene of interest.
Even though the tree has many other different species entered in to it the tree does still hold consistent with what is found through the phylogenetic trees based off of the protein sequences within the Protein Phylogeny section.
Even though the tree has many other different species entered in to it the tree does still hold consistent with what is found through the phylogenetic trees based off of the protein sequences within the Protein Phylogeny section.
