This web page was produced as an assignment for Genetics 677, and undergraduate course at UW-Madison.
MEME DNA Motifs

MEME (Multiple Em for Motif Elicitation) is a motif-detecting software programs which can detect short, reoccurring DNA patterns that might have biological fucntions. These motifs usually indicate sequence specific binding sites for proteins that are involved in biological processes in the RNA level (ribosome binding, mRNA splicing, transcription). Instead of using the regular MEME program MEME-ChIP was used as it allows for larger DNA sequences to be searched for motifs. The full KCNQ1 gene was used to detect the DNA motifs in the gene in the MEME program.
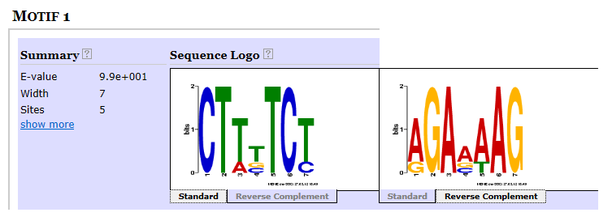

The sequence on the left shows the actual motif sequence while the sequence on the right shows the complementary sequence to the motif sequence.
Again the actual motif sequence is on the left while the complementary sequence is on the right.
Again the actual motif sequence is on the left while the complementary sequence is on the right.
Gene Motifs
Gene Found Motifs
Just like shown above the list below is a list of motifs that were found using another Motif search through GenomeNet. Through this search 14 different motifs were found.
EGF_1: EGF-like domain signiture 1
Prosite: PS00022
J_ACTX: Janus-faced atracotoxin (J-ACTX) family signature
Prosite: PS60020
INTEGRIN_BETA: Integrins beta chain cysteine-rich domain signature
Prosite: PS00243
CTCK_1: C-terminal cysteine knot signature
Prosite: PS01185
ANAPHYLATOXIN_1: Anaphylatoxin domain signature
Prosite: PS01177
AGOUTI_1: Agouti domain signature
Prosite: PS60024
IGFBP_N_1: Insulin-like growth factor-binding protein (IGFBP) N-terminal domain signature
Prosite: PS00222
THIOLASE_3: Thiolases active site
Prosite: PS00099
TUBULIN: Tubulin subunits alpha, beta, and gamma signature
Prosite: PS00227
HYDROPHOBIN: Fungal hydrophobins signature
Prosite: PS00956
VWFC_1: VWFC domain signature
Prosite: PS01208
4FE4S_FER_1: 4Fe-4S ferredoxin type iron-sulfur binding region signature
Prosite: PS00198
2FE2S_FER_1: 2Fe-2S ferredoxin type iron-sulfur binding region signature
Prosite: PS00197
DEFENSIN: Mammalian defensins signature
EGF_1: EGF-like domain signiture 1
Prosite: PS00022
J_ACTX: Janus-faced atracotoxin (J-ACTX) family signature
Prosite: PS60020
INTEGRIN_BETA: Integrins beta chain cysteine-rich domain signature
Prosite: PS00243
CTCK_1: C-terminal cysteine knot signature
Prosite: PS01185
ANAPHYLATOXIN_1: Anaphylatoxin domain signature
Prosite: PS01177
AGOUTI_1: Agouti domain signature
Prosite: PS60024
IGFBP_N_1: Insulin-like growth factor-binding protein (IGFBP) N-terminal domain signature
Prosite: PS00222
THIOLASE_3: Thiolases active site
Prosite: PS00099
TUBULIN: Tubulin subunits alpha, beta, and gamma signature
Prosite: PS00227
HYDROPHOBIN: Fungal hydrophobins signature
Prosite: PS00956
VWFC_1: VWFC domain signature
Prosite: PS01208
4FE4S_FER_1: 4Fe-4S ferredoxin type iron-sulfur binding region signature
Prosite: PS00198
2FE2S_FER_1: 2Fe-2S ferredoxin type iron-sulfur binding region signature
Prosite: PS00197
DEFENSIN: Mammalian defensins signature
Analysis
By going through all the domains of the KCNQ1 DNA sequence many conserved regions were found. The GenomeNet database found many sites that show different components of KCNQ1 that are important in its function. Potential mutations could be accuring within these conserved areas and could be topics for future research to further our understanding as to how KCNQ1 works and functions within the cells. It may be found that certain signatures are only functioning in certain cells and others function in other cells.

