This web page was produced as an assignment for Genetics 677, and undergraduate course at UW-Madison.
Protein Phylogeny
Phylogenetic Trees using Phylogeny.fr
Phylogeny.fr phylogenetic tree using MUSCLE Alignment
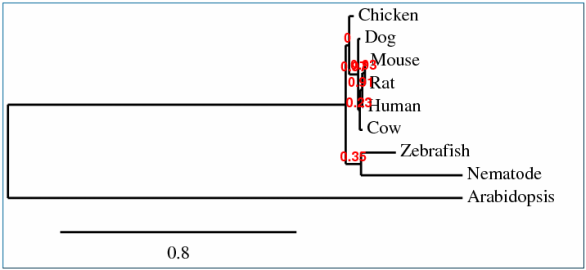
This phylogenetic tree incorporates the phylogram approach of making a tree that uses the length of its branches to display the amount of time that has passed between the different divergence points. The longer the branch is the more time that has passed since the species have splits.
Phylogeny.fr phylogenetic tree using MUSCLE Alignment
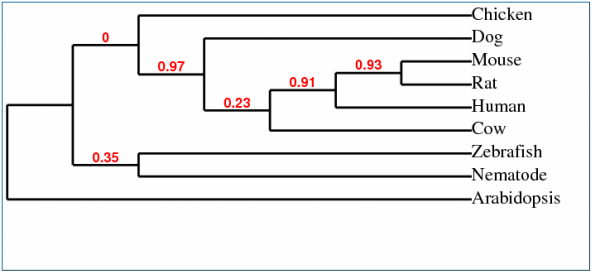
Unlike the phylogenetic tree shown above this one is an example of a cladogram which does not show the time since the species has split. This makes it easier to see the divergence between species that have split more recently in history.
Phylogeny.fr phylogenetic tree using ClustalW Alignment
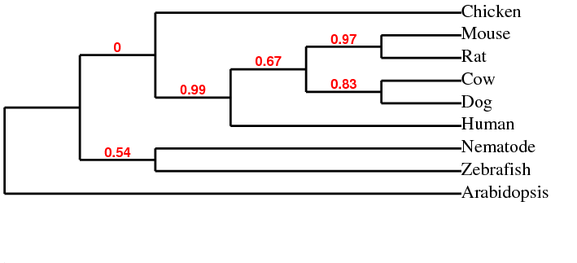
Phylogeny.fr phylogenetic tree using T-Coffee Alignment
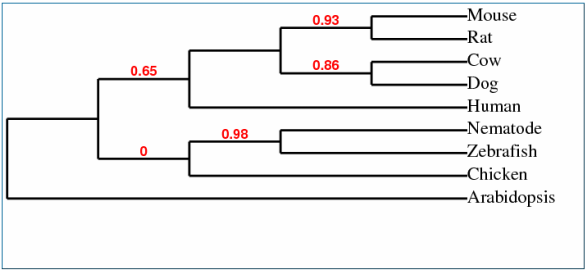
_Phylogenetic Tree using ClustalW alignment in GeneBee
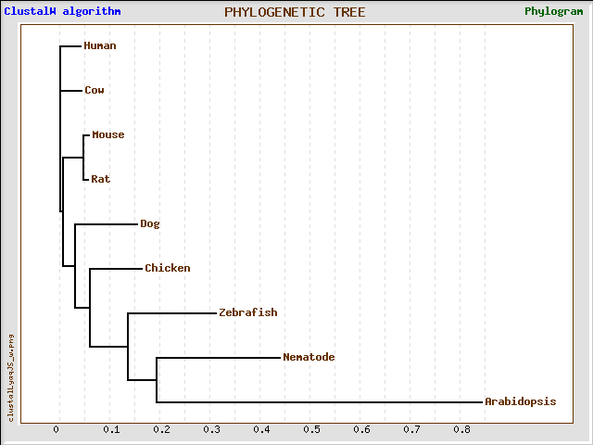
_This phylogenetic tree was generated by the GeneBee service using the ClustalW protein alignment results.
Phylogenetic Tree using Muscle alignment in GeneBee
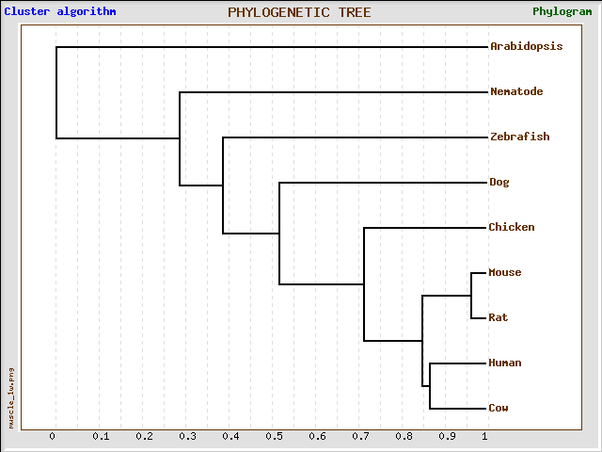
This phylogenetic tree was generated by the GeneBee service using the MUSCLE protein alignment results.
Analysis
Even though it was the same sequences used there were slight variations of how the trees ended as to where the branch points were for the mammalian species. It was interesting that in some trees the cow was shown to be recently divergered with the human while other trees show the human by itself and the cow closely related with dog. Perhaps there are certain aspects of the proteins that share more conservation between cows and humans while others share more between cows and dogs. Perhaps another look at the sequences using shorter se


