This web page was produced as an assignment for Genetics 677, and undergraduate course at UW-Madison.
Protein Domains
_To determine the domains found in the KCNQ1 protein sequence SMART and Pfam, two online programs, were used. Both programs returned similar domain results but SMART returned a little more information as well as an extra significant domain over what Pfam returned. The 3D structures were available though through the Pfam website and not through SMART which allowed another look as to what the protein domains may look like. The results obtained from each program and the interaction maps formed are discussed below.
Pfam
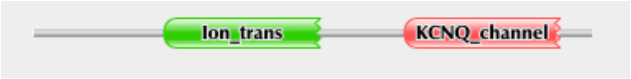
_Using the protein sequence for the KCNQ1 protein, a search through the Pfam database resulted in the protein map shown above. The two domains shown are two that the site deemed to be significant. The green domain shown represents Ion-trans, a Ion transport protein found about a third of the way into the protein from the N-terminus. Pfam found this domain to begin at position 158 and end at position 346, with an e-value of 3.5e-22. The red domain represents the KCNQ-channel domain, this domain turns out to be the KCNQ voltage-gated potassium channel. Pfam found this KCNQ_channel domain to begin at position 451 and end at position 634, with an e-value of 4.4e-64. Besides the two significant domains shown, Pfam also found another domain but deemed it insignificant and was not shown in the protein map. All of the significant domain matches obtained from the search sequence were done so using Pfam-A under the default settings.
SMART
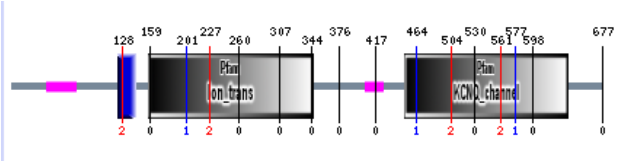
Using the same protein sequence as used in the Pfam, a search through the SMART database returned the protein map shown above. SMART found the same Ion-trans domain as seen above in the Pfam search, beginning at position 158 and ending at position 346, with an e-value of 1.00e-19. SMART also found the KCNQ-channel domain from the Pfam search, beginning at position 451 and ending at position 639, with an e-value of 3.8e-61. This domain is described to be involved in shaping the action potential, and in neuronal excitability and plasticity. [3] SMART also found an transmembrane region beginning at position at position 123 and ending at position 145. This region is represented in blue in the above image, an e-value was not giving for this domain. Two other insignificant protein domains are shown in purple. These two were dismissed as they have not shown to be important in the use of the protein at this point in time.
3D Protein Structures
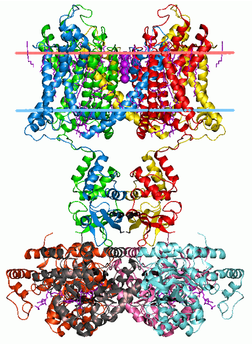
The figure to the left shows the 3D structure of the two main domains found in the KCNQ1 protein. Since both of the domains are for the ion channels they both have similar 3D structures which is why only one image is shown for the two domains discussed above from the SMART and Pfam search results, Ion_trans and KCNQ_channel. [4]
Analysis
Even though there were only two (three if you look at the SMART entry) domains within the KCNQ1 protein does not mean that there is not a less importance to the protein. Those two domains play a critical role in keeping the heart beating and one small mutation in either of the domains could spell huge problems for the individual.
It is also these domains that lead me to new ideas as to what could be going wrong to cause the symptoms of Long QT Syndrome. The domains are highly interactive with many proteins and it was these proteins that I choose to look at within my experiment to see if there were other things that could lead to the same diagnosis of Long QT Syndrome.
It is also these domains that lead me to new ideas as to what could be going wrong to cause the symptoms of Long QT Syndrome. The domains are highly interactive with many proteins and it was these proteins that I choose to look at within my experiment to see if there were other things that could lead to the same diagnosis of Long QT Syndrome.